ID.9 |
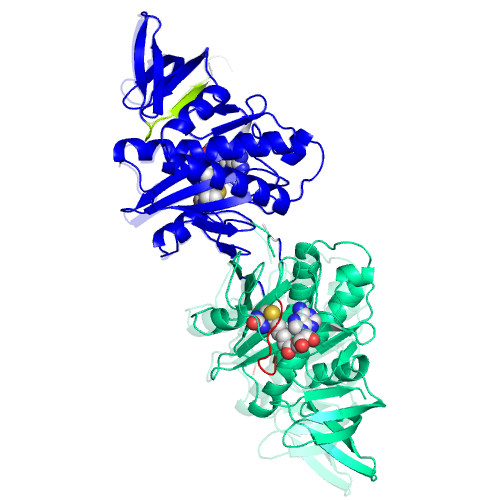
235AA LONG HYPOTHETICAL BIOTIN--[ACETYL-COA-CARBOXYLASE] LIGASE |
|
|
|
Function |
EC*1 |
CSA distance*2 |
6.3.4.15 |
2.6 |
|
*1 Enzyme commission number. |
Ligand |
PDB*1 |
Full name |
2xADP,2xBTN |
2x(ADENOSINE-5'-DIPHOSPHATE),2x(BIOTIN) |
|
*1 Ligand name designated by the PDB identifiers. |
Segments |
Component No. |
Fixed*1 |
Moving*1 |
Motion type |
Ligand binding |
Coupled motion type |
1 |
D1(3A-4A,10A-12A,5B-110B,112B-233B) |
D2(5A-9A,13A-48A,54A-110A,112A-235A,1B-4B) |
Domain |
Independent |
|
2 |
D1 |
L3(227B-233B) |
Local |
Independent |
|
3 |
D2 |
L4(48A-54A) |
Local |
Coupled |
Closure |
|
*1 The location of the fixed and moving segments indicated by the residue number assigned in the ligand-bound form. The background color of characters indicates the corresponding segment in the structure. The colored segments not described in the Table are: 1) a part of component in which the motion is small (< 1.0 A), or, 2) a part of a protomer of homodimers, for which a corresponding part of the other protomer is shown in the Table. |
Displacement and disorder |
Component No. |
RMSD*1 |
Displacement*2 |
Disorder-order transition*3 |
Disorder residue*4 |
Helix-Coil*5 |
1 |
1.4 |
||||
2 |
5.5 |
||||
3 |
Yes |
+5 |
|
*1 The root-mean-square displacement of a component of motion calculated for the domain motions. |
Crystal environment |
Component No. |
Open state*1 |
Crystal packing*2 |
1 |
Free |
Coupled |
2 |
Bound |
Coupled |
3 |
|
*1 Distinction between the open state and the closed state required for examining the influence of the crystal environment. |